I love GPU programming for precisely this: it forces and enables you to think about a solution in a non-linear fashion in more than one sense of the word.
The Problem
Given a set , output a set
In other words, pretend our array is cyclical, and we want all partial sums of the array elements, in order, starting from element i, ending with i + j. (We will be looping through the end, since the array is cyclical.)
In the bioinformatics course, this happens to be a toy problem of generating a cyclo-spectrum from the weights of a cyclic peptide.
So for a peptide with individual amino-acid masses [57; 71; 113; 113; 128], the first 3 iterations will look like:
| 57 | 71 | 113 | 113 | 128 |
| 57 + 71 | 71 + 113 | 113 + 113 | 113 + 128 | 128 + 57 |
| 57 + 71 + 113 | 71 + 113 + 113 | 113 + 113 + 128 | 113 + 128 + 57 | 128 + 57 + 71 |
Code
On my GitHub.
Sequential Solutions
The above table immediately hints at a brute-force as well as an optimized solution. The brute-force solution is :
// brute force O(n^3) solution
let cyclospec (peptide : int seq) =
let len = peptide.Count()
let pepArr = peptide |> Seq.toArray
let mutable parsum = 0
(seq {
yield 0
yield pepArr |> Array.sum
for i = 0 to len - 2 do
yield! [0..len-1]
|> Seq.map
(fun j ->
//F# 4.0 Feature!
parsum <- 0
for ind = j to j + i do
parsum <- parsum + pepArr.[if ind >= len then ind - len else ind]
parsum
)
}) |> Seq.sort |> Seq.toArray
Note the F# 4.0 feature: a variable mutated within a closure.
This will work, but slowly. To significantly speed it up we only need to notice that there is no need to recompute values that have been precomputed already over, and over, and over again. So, memoization yields an solution:
let cyclospecOpt (peptide : int seq) =
let len = peptide.Count()
let pepArr = peptide |> Seq.toArray
let output = Array.zeroCreate ((len - 1) * len)
for i = 0 to len - 2 do
[0..len-1]
|> Seq.iter
(fun j ->
let ind = i + j
output.[i * len + j] <-
if i = 0 then pepArr.[j]
else output.[(i - 1) * len + j] + pepArr.[if ind >= len then ind - len else ind]
)
seq {yield 0; yield pepArr |> Array.sum; yield! output} |> Seq.sort |> Seq.toArray
Ok, so far so good.
Multi-dimensional thinking with CUDA
So how do we look at this differently with CUDA? I believe solving this problem shows how data, for lack of a better term, fuses with time (represented by threads). If we take a closer look at the table above, we’ll see that the solution can be represented in a two-dimensional grid. For instance, at grid location (5, 3), we have a solution element that is constructed of , thus it’s a grid X of dimensions n x (n-1)( where
is the sum, starting from i-th element of the input set, and ranging over j elements.
This data structure maps easily into CUDA with its 3D thread model (this is the element of time/data fusion I was talking about).
Since we now have ALEA.Cuda, we can do this without leaving F#, and, even better, we can script it:
[<Kernel; ReflectedDefinition>]
let cyclospecKernel (arr : deviceptr<int>) (len : int) (out : deviceptr<int>) =
let ind = blockIdx.x * blockDim.x + threadIdx.x
let lenElem = blockIdx.y * blockDim.y + threadIdx.y
if ind < len && lenElem < len - 1 then
let mutable parsum = 0
for i = 0 to lenElem do
let idx = ind + i
parsum <- parsum + arr.[if idx >= len then idx - len else idx]
out.[lenElem * len + ind] <- parsum
And we invoke it:
let cyclospecGpu (peptide : int []) =
let blockSize = dim3(16, 16)
let gridSize = dim3(divup peptide.Length blockSize.x, divup peptide.Length blockSize.y)
let lp = LaunchParam(gridSize, blockSize)
use dPeptide = worker.Malloc(peptide)
use dOutput : DeviceMemory<int> = worker.Malloc(peptide.Length * (peptide.Length - 1))
worker.Launch <@cyclospecKernel @> lp dPeptide.Ptr peptide.Length dOutput.Ptr
let output = dOutput.Gather()
seq{yield 0; yield! output; yield peptide |> Seq.sum} |> Seq.toArray |> Array.sort
Each kernel thread is computing its own element in the grid, which is flattened into a solution. Time has indeed merged with data.
But what about our optimization? The easy way to implement it is to streamline invocations of a kernel, where each consecutive run will compute the new sum using the results of previous kernel invocations. There are variations on this theme, but it lacks the aesthetic value of the “brute force” CUDA solution:
[[]
let cyclospecOptKernel (arr : deviceptr) (len : int) (out : deviceptr) (lenElem : int)=
let ind = blockIdx.x * blockDim.x + threadIdx.x
if ind < len then
let idx = ind + lenElem
out.[lenElem * len + ind] = len then idx – len else idx]
Invocation:
let cyclospecOptGpu (peptide : int []) =
let blockSize = 256
let gridSize = divup peptide.Length blockSize
let lp = LaunchParam(gridSize, blockSize)
use dPeptide = worker.Malloc(peptide)
use dOutput : DeviceMemory<int> = worker.Malloc(peptide.Length * (peptide.Length - 1))
for i = 0 to peptide.Length - 2 do
worker.Launch <@cyclospecOptKernel @> lp dPeptide.Ptr peptide.Length dOutput.Ptr i
let output = dOutput.Gather()
seq{yield 0; yield! output; yield peptide |> Seq.sum} |> Seq.toArray |> Array.sort
So, how did we do? (Here comes the mandatory chart):
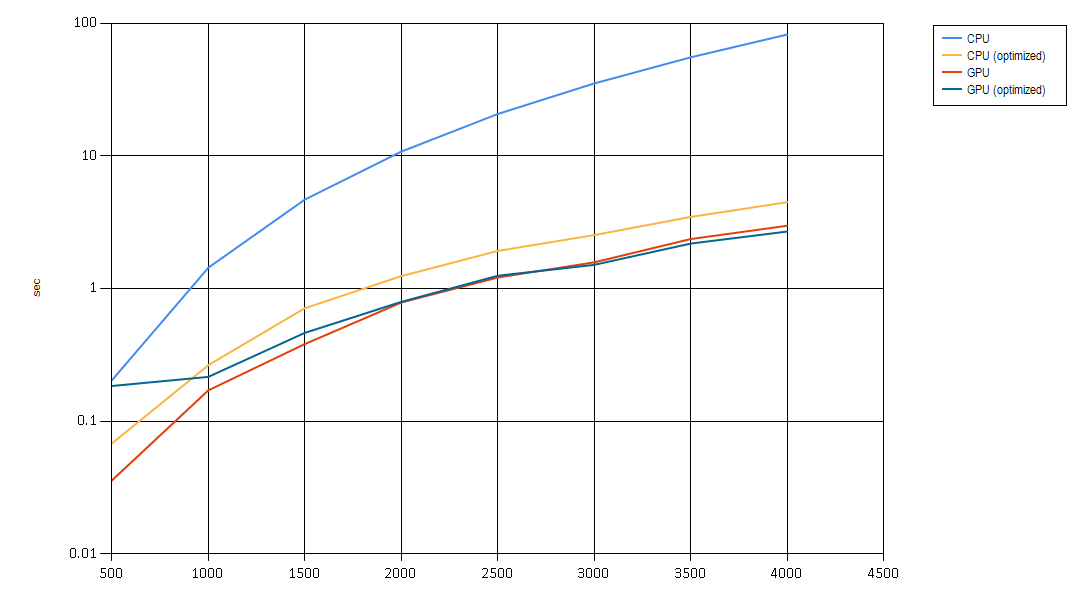
And so:
- A little algorithm is a beautiful thing
- A little CUDA is an even more beautiful thing
- CUDA optimization performs no worse and at larger sizes of the input starts performing better, at some point massive amounts of computation slow us down even in parallel. See #1.
- And what is the GPU algorithm performance in O notation? It’s definitely non-linear, is it possible to estimate analytically?
Here is another chart that only compares optimized algorithms. It’s not a semi-log chart for easier visualization:
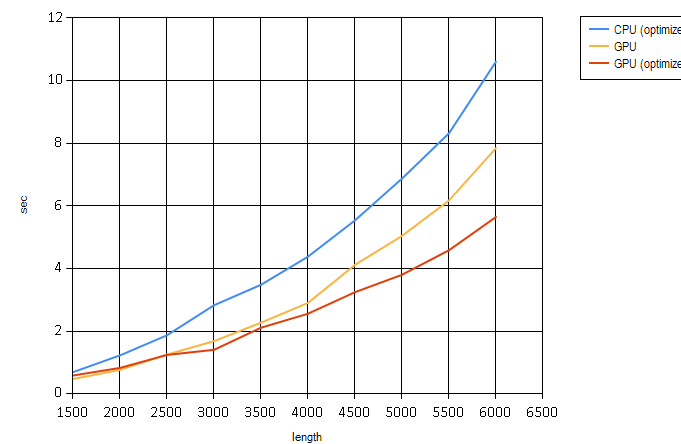
One thought on “Non-linear Thinking with CUDA.”